-Search query
-Search result
Showing 1 - 50 of 108 items for (author: fujita & j)

EMDB-36724: 
Structure of the SARS-CoV-2 XBB.1.5 spike glycoprotein (closed state 1)
Method: single particle / : Yajima H, Anraku Y, Kita S, Kimura K, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-36726: 
Structure of the SARS-CoV-2 XBB.1.5 spike glycoprotein (closed state 2)
Method: single particle / : Yajima H, Anraku Y, Kita S, Kimura K, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-36727: 
Structure of SARS-CoV-2 XBB.1.5 spike glycoprotein in complex with ACE2 (1-up state)
Method: single particle / : Yajima H, Anraku Y, Kita S, Kimura K, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-36728: 
Structure of SARS-CoV-2 XBB.1.5 spike glycoprotein in complex with ACE2 (2-up state)
Method: single particle / : Yajima H, Anraku Y, Kita S, Kimura K, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-36729: 
Structure of SARS-CoV-2 XBB.1.5 spike RBD in complex with ACE2
Method: single particle / : Yajima H, Anraku Y, Kita S, Kimura K, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-36251: 
RNA polymerase II elongation complex bound with Elf1, Spt4/5 and foreign DNA, stalled at SHL(-1) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

EMDB-36252: 
RNA polymerase II elongation complex containing 40 bp upstream DNA loop, stalled at SHL(-1) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

EMDB-36253: 
RNA polymerase II elongation complex containing 60 bp upstream DNA loop, stalled at SHL(-1) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

EMDB-37848: 
RNA polymerase II elongation complex bound with Elf1, Spt4/5 and foreign DNA, stalled at SHL(0) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

PDB-8jh2: 
RNA polymerase II elongation complex bound with Elf1, Spt4/5 and foreign DNA, stalled at SHL(-1) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

PDB-8jh3: 
RNA polymerase II elongation complex containing 40 bp upstream DNA loop, stalled at SHL(-1) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

PDB-8jh4: 
RNA polymerase II elongation complex containing 60 bp upstream DNA loop, stalled at SHL(-1) of the nucleosome
Method: single particle / : Akatsu M, Fujita R, Ogasawara M, Ehara H, Kujirai T, Takizawa Y, Sekine S, Kurumizaka H

EMDB-33785: 
Cryo-EM structure of the histamine-bound histamine H4 receptor and Gq complex
Method: single particle / : Im D, Iwata S, Asada H

EMDB-33786: 
Cryo-EM structure of the imetit-bound histamine H4 receptor and Gq complex
Method: single particle / : Im D, Iwata S, Asada H

EMDB-34871: 
Cryo-EM structure of biparatopic antibody Bp109-92 in complex with TNFR2
Method: single particle / : Akiba H, Fujita J, Ise T, Nishiyama K, Miyata T, Kato T, Namba K, Ohno H, Kamada H, Nagata S, Tsumoto K

PDB-8hlb: 
Cryo-EM structure of biparatopic antibody Bp109-92 in complex with TNFR2
Method: single particle / : Akiba H, Fujita J, Ise T, Nishiyama K, Miyata T, Kato T, Namba K, Ohno H, Kamada H, Nagata S, Tsumoto K

EMDB-36048: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36049: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36050: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36051: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36052: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36053: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36055: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7t: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7u: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7v: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7w: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7x: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7y: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j80: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S
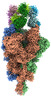
EMDB-33972: 
Cryo-EM structure of the SARS-CoV-2 spike protein (3-up RBD) bound to neutralizing antibody CSW1-1805
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
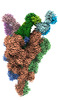
EMDB-33973: 
Cryo-EM structure of the SARS-CoV-2 spike protein (1-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
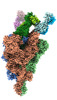
EMDB-33974: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
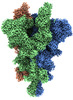
EMDB-33975: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) with C480A mutation
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y

EMDB-34429: 
Cryo-EM structure of KpFtsZ-monobody double helical tube
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

PDB-8h1o: 
Cryo-EM structure of KpFtsZ-monobody double helical tube
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

EMDB-35344: 
Cryo-EM structure of KpFtsZ single filament
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hibino K, Konishi T, Kato Y, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

PDB-8ibn: 
Cryo-EM structure of KpFtsZ single filament
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hibino K, Konishi T, Kato Y, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

EMDB-29665: 
Cryo-EM structure of Nav1.7 with CBD
Method: single particle / : Fan X, Huang J, Yan N

EMDB-35622: 
SARS-CoV-2 XBB.1 spike glycoprotein (closed-1 state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-35623: 
SARS-CoV-2 XBB.1 spike glycoprotein (closed-2 state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-35624: 
SARS-CoV-2 XBB.1 spike glycoprotein in complex with ACE2 (1-up state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-35626: 
SARS-CoV-2 XBB.1 spike glycoprotein in complex with ACE2 focused on RBD-ACE2 interface
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

PDB-8ios: 
Structure of the SARS-CoV-2 XBB.1 spike glycoprotein (closed-1 state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

PDB-8iot: 
Structure of the SARS-CoV-2 XBB.1 spike glycoprotein (closed-2 state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

PDB-8iou: 
Structure of SARS-CoV-2 XBB.1 spike glycoprotein in complex with ACE2 (1-up state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

PDB-8iov: 
Structure of SARS-CoV-2 XBB.1 spike RBD in complex with ACE2
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-35625: 
SARS-CoV-2 XBB.1 spike glycoprotein in complex with ACE2 (2-up state)
Method: single particle / : Anraku Y, Kita S, Yajima H, Sasaki J, Sasaki-Tabata K, Maenaka K, Hashiguchi T

EMDB-34016: 
Lloviu cuevavirus nucleoprotein-RNA complex
Method: single particle / : Hu SF, Fujita-Fujiharu Y, Sugita Y, Wendt L, Muramoto Y, Nakano M, Hoenen T, Noda T
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
